Recently, the Biotechnology Research Institute (BRI) of the Chinese Academy of Agricultural Sciences (CAAS) made new progress in the study of cotton fiber domestication and breeding selection. The research team identified that GhTT2-A07, a gene encoding an MYB transcription factor, regulates both fiber color and quality simultaneously. Natural variations in the gene’s promoter region were directionally selected during breeding, serving as a key genetic factor in the development of modern white, high-quality cotton. These findings were published in the internationally renowned academic journal The Plant Journal.
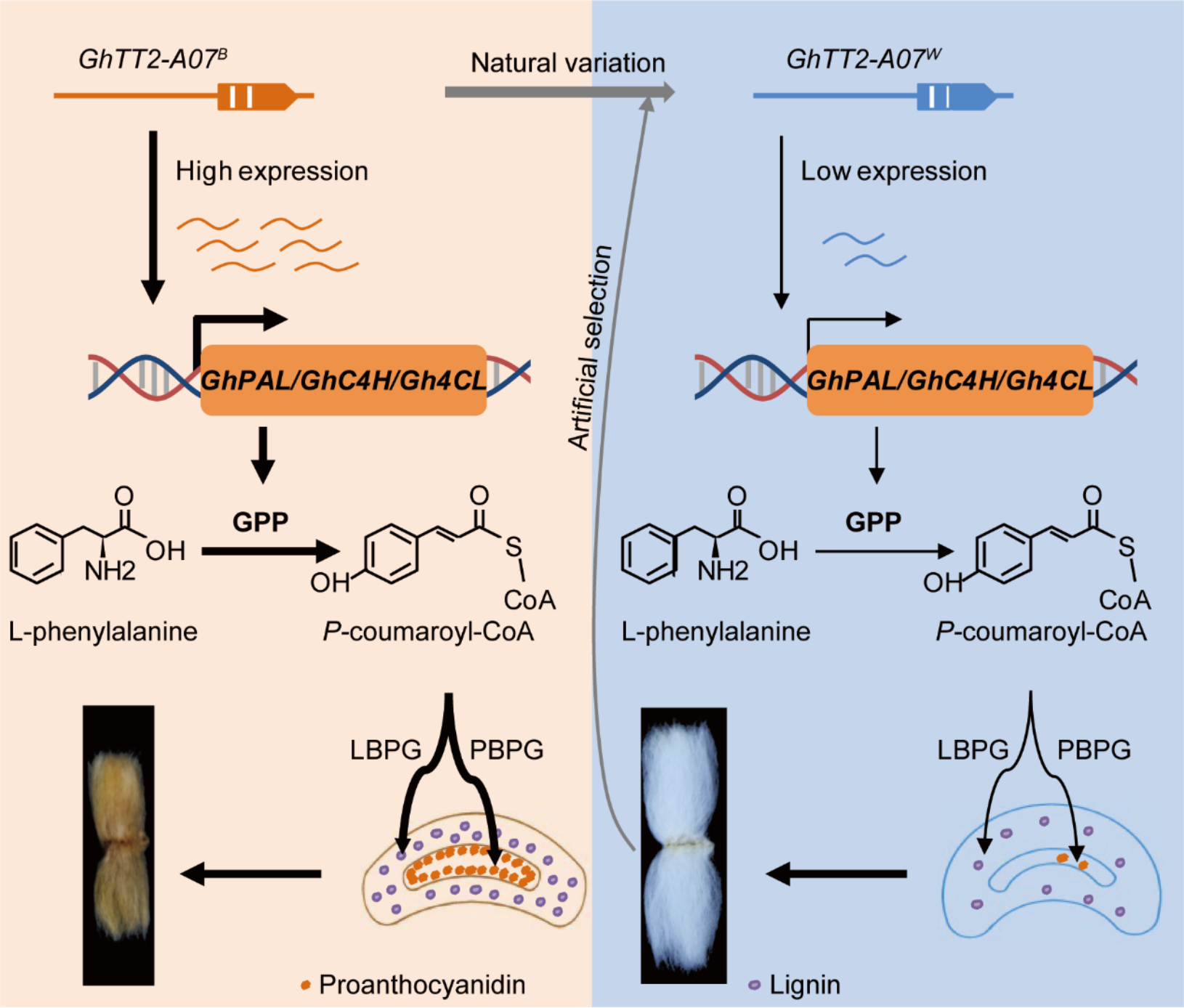
Figure. Regulatory model of GhTT2-A07 in cotton fiber development and its evolutionary selection in modern breeding.
Cotton is an important economic crop with a long history of domestication and breeding improvement. Currently widely cultivated upland cotton (Gossypium hirsutum) and island cotton (G. barbadense); both are allotetraploids, derived from the hybridization and genome doubling of diploid ancestors. While many wild tetraploid cotton species naturally produce colored fibers, artificial selection during domestication gradually favored white-fibered varieties as the dominant cultivated type. Previous studies have shown that naturally colored cotton fibers generally have inferior agronomic qualities. Although, modern white cotton cultivars evolved from these ancestral colored varieties, but the molecular mechanisms behind this transition remained unclear.
To unravel the regulatory networks governing cotton fiber domestication, the researchers used ZXM1, a brown-fibered cotton line, as experimental material. Through map-based cloning, they successfully identified GhTT2-A07—an R2R3-MYB transcription factor-encoding gene that controls both fiber pigmentation and length. In brown cotton fibers, GhTT2-A07 is specifically and highly expressed during development, activating downstream genes in the phenylpropanoid pathway, including GhPALs, GhC4Hs, and Gh4CLs. This activation triggers the accumulation of lignin and proanthocyanidins in fiber cells. These secondary metabolites give fibers natural brown color, but interferes with the oriented assembly of cellulose microfibrils in the cell wall, inhibiting fiber elongation and reducing fiber strength.
During the historical process of cotton breeding, natural variations in the GhTT2-A07 promoter region significantly down-regulated the gene’s expression during fiber development. This genetic change effectively reduced pigment biosynthesis while promoting fiber cell elongation and development. Haplotype analysis further confirmed that the low-expression haplotype of GhTT2-A07 underwent strong positive selection during cotton evolution and was ultimately fixed in modern high-quality white cotton germplasm. This study not only provides critical insights into the molecular regulation of cotton fiber development but also deepens understanding of the coordinated evolutionary trajectory of fiber color and quality during cotton domestication and breeding.
Co-first authors of the paper are He Haiyan, a PhD candidate, Zhang Jilong, a graduated master’s student, and Zhou Qi, a postdoctoral researcher, all from BRI. Professor Liang Chengzhen is the corresponding author. Wang Hongzhe and Wen Ting from Xinjiang Guoxin Seed Industry provided generous material support, while bioinformatics analyses were supported by Dr. Gao Qiang from Qi Biodesign and Professor Ou Shujun from The Ohio State University. This research was funded by the National Key Research and Development Program of China, the National Natural Science Foundation of China, and other projects.
Original article: https://doi.org/10.1111/tpj.70863 |